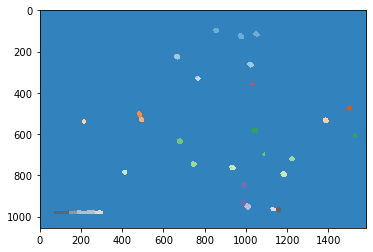
I have the following image. I was able to use watershed to detect all the particles using the code below.
I was able to use watershed to detect all the particles using the code below.
However, now I need to calculate the size of each particles in the figure and if I use the "labels" image, for some reasons I am not capable of using the function cv2.findContours.
Anyone willing to share some ideas? If you propose some code, please include explanation because I am a beginner. :)
Many thanks!
import numpy as np
import cv2
import matplotlib.pyplot as plt
from scipy import ndimage as ndi
from skimage.morphology import watershed
from skimage.feature import peak_local_max
#-------------------------------------------------------------------------------------------#
# IMAGE PRETREATMENT
img = cv2.imread('Test images/TEM of nanoparticles/NP good 0010.tif')
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
Gaussian_Blur = cv2.GaussianBlur(gray,(21, 21), cv2.BORDER_DEFAULT)
# Use fixed threshold to mask black areas
_, thresh = cv2.threshold(Gaussian_Blur, 90, 255, cv2.THRESH_BINARY_INV) # _ = 30
# Morphological closing to close holes inside particles; opening to get rid of noise
img_mop1 = cv2.morphologyEx(thresh, cv2.MORPH_CLOSE, cv2.getStructuringElement(cv2.MORPH_ELLIPSE, (7, 7)))
img_mop = cv2.morphologyEx(img_mop1, cv2.MORPH_OPEN, cv2.getStructuringElement(cv2.MORPH_ELLIPSE, (15, 15)))
tiled_h = np.hstack((img_mop1, img_mop)) # stack images side-by-side
plt.figure('Pretreatment')
plt.subplot(2, 2, 1) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('Gray')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(gray, cmap='gray')
plt.subplot(2, 2, 2) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('Gaussian_Blur')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(Gaussian_Blur, cmap='gray')
plt.subplot(2, 2, 3) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('Thresh')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(thresh, cmap='gray')
plt.subplot(2, 2, 4) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('img_mop')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(img_mop, cmap='gray')
#-------------------------------------------------------------------------------------------#
# WTERSHED WITH SKIMAGE
# Now we want to separate the two objects in image
# Generate the markers as local maxima of the distance to the background
distance = ndi.distance_transform_edt(img_mop) # Calculates distance of pixels from background
#Find peaks in an image as coordinate list or boolean mask.
local_maxi = peak_local_max(distance, indices=False, footprint=np.ones((3, 3)), labels=img_mop)
# indices: if True, the output will be an array representing peak coordinates. If False, the output will be a boolean
# array shaped as image.shape with peaks present at True elements.
# If footprint == 1 represents the local region within which to search for peaks at every point in image.
# labels: if provided, each unique region labels == value represents a unique region to search for peaks. Zero is
# reserved for background.
# returns an array of boolean with True on max points
print('local_maxi lenght: ', local_maxi.shape)
print('local_maxi: ', local_maxi[0])
markers = ndi.label(local_maxi)[0]
print('markers lenght: ', markers.shape)
print('markers: ', markers[0])
labels = watershed(-distance, markers, mask=img_mop)
print('labels lenght: ', labels.shape)
print('labels: ', labels[0])
plt.figure('Processing')
plt.subplot(2, 2, 1) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('Distance trans')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(distance, cmap='gray')
plt.subplot(2, 2, 2) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('local_maxi')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(local_maxi, cmap='gray')
plt.subplot(2, 2, 3) # Figure two has subplots 2 raw, 2 columns, and this is plot 1
plt.gca().set_title('markers')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(markers, cmap='gray')
plt.figure('Watershed')
plt.gca().set_title('Watershed')
plt.xticks([]), plt.yticks([]) # To hide axes
plt.imshow(labels, cmap='gray')
plt.show()
#-------------------------------------------------------------------------------------------#
# DATA ANALYSIS ---- WORK IN PROGRESS
cnts, _ = cv2.findContours(labels, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_NONE)
img = cv2.drawContours(img, cnts, -1, (0, 255, 255), 2) # To print all contours
cv2.imshow('Contours', cv2.resize(img, dsize=(0, 0), fx=0.3, fy=0.3))
print('\nCnts length: ', len(cnts), '\n') # 11 objects (10 nanoparticles + scale barr)
# Divide the cnts array into scalebar and nanoparticles
# Get bounding rectangles for the scale and the particles from detailed contour determine on line 32.
# cv2.boundingRect() outputs: x, y of starting point (top left corner), and width and height of rectangle.
# Find contours. For more info see: https://opencv-python-tutroals.readthedocs.io/en/latest/py_tutorials/py_imgproc/py_contours/py_contour_features/py_contour_features.html
# cv2.contourArea() outputs the area of each detailed contour, does not work on rectangle generated by cv2.boundingRect.
thr_size = 5000
for cnt in cnts:
if cv2.contourArea(cnt) > thr_size:
scale = [cv2.boundingRect(cnt)] # returns x, y, w, h
img = cv2.rectangle(img, (scale[0][0], scale[0][1]), (scale[0][0] + scale[0][2], scale[0][1] + scale[0][3]), (255, 255, 0), 2)
print('Scale is: ', scale) #only one box (object) = scalebar
print("scale[0][1] is scalebar's width of {} pixels".format(scale[0][2]), '\n')
# 8. MINIMUM ENCLOSING CIRCLE
i = 1
for cnt in cnts:
if cv2.contourArea(cnt) < thr_size:
# Find min enclosing circle and get xy of centre
(x, y), radius = cv2.minEnclosingCircle(cnt)
center = (int(x), int(y))
# Get radius average method
#rx, ry, w, h = cv2.boundingRect(cnt)
#radius = int((((w+h)/2))*1.5)
img = cv2.circle(img, center, radius, (255, 0, 255), 3)
cv2.putText(img, str(i), (int(x), int(y)-20), cv2.FONT_HERSHEY_COMPLEX, 1, (0, 255, 0), 2)
print('Particle ' + str(i) + ' | Horizontal diameter: ' + '{:.2f}'.format((radius/ scale[0][2] * 200)*2) + ' nm')
i=i+1
cv2.imshow('img', cv2.resize(img, dsize=(0, 0), fx=0.3, fy=0.3))